London Calling was in a new venue versus the previous year, having outgrown the sleek Milbank Tower and moved into much more spacious confines of Old Billingsgate. Milbank Tower was on the edge of a posh, leafy residential neighborhood and had long views of the city from the top floor (and interminably long waits for the lifts to go to-and-from those views!). Float down the Thames just shy of five kilometers and you'll arrive at Old Billingsgate, a squat one-time fishmarket in the heart of The City of London, the financial hub. So if I wanted to force a metaphor, both the conference and company are drifting down from the clouds and trees and now getting down to business
Clive opened his talk with a warning, which the audience chuckled at, that his talk would be restrained versus previous year's talk. Then the delegates proceeded to be shocked as he largely stuck to that prediction. Whereas last year's commotion brought out the ideas of Zumbador and SmidgeION, this year lacked any extravagant new visions. It's also the case that ONT is no longer saving a year's worth of vision for a single date in May; Clive's online updates in October and March unveiled Flongle and the FPGA compute dongle respectively. There's also the issue, as will be detailed, of catching up with prior pronouncements. ONT hasn't been as bad at this as is sometimes portrayed -- the past three years have seen a steady progression in the platform -- but there is still a lot of catching up on major systems.
MinION
There wasn't a lot of talk about MinION, but it didn't go ignored.Two major changes in flowcells are projected for this summer which will have multiple benefits. First, "no priming" flowcells are undergoing final stability testing, eliminating a time-consuming step which flusters many novices (but not me anymore!). Second, flowcells will be moved to shipping at ambient temperature, wrapped in wool. Clive expressed both his hatred of having a cold chain as well as a quirky vision of users knitting their packing material into jumpers. One tweeter even suggested trading wool for flowcells!
The stability data shown for the ambient shipping would also seem to support longer warranties than the current 10-day one ONT offers, though no mention of such an extension was made. Certainly many users (including myself) report aged flowcells performing well.
Clive remains bullish on the power of FPGAs to accelerate basecalling. ONT is targeting fall for alpha release of a dongle to place FPGAs between the MinION and the controlling computer. The device doesn't yet have a name (Clive confessed to a bit of block on naming), leading to some tweeters making suggestions such as PortION (Mick Watson) and ORCA (online realtime caller, from Nicolas Coutin). Amanda Warr pointed to a link for all the English words which end with "ion"
.

PromethION
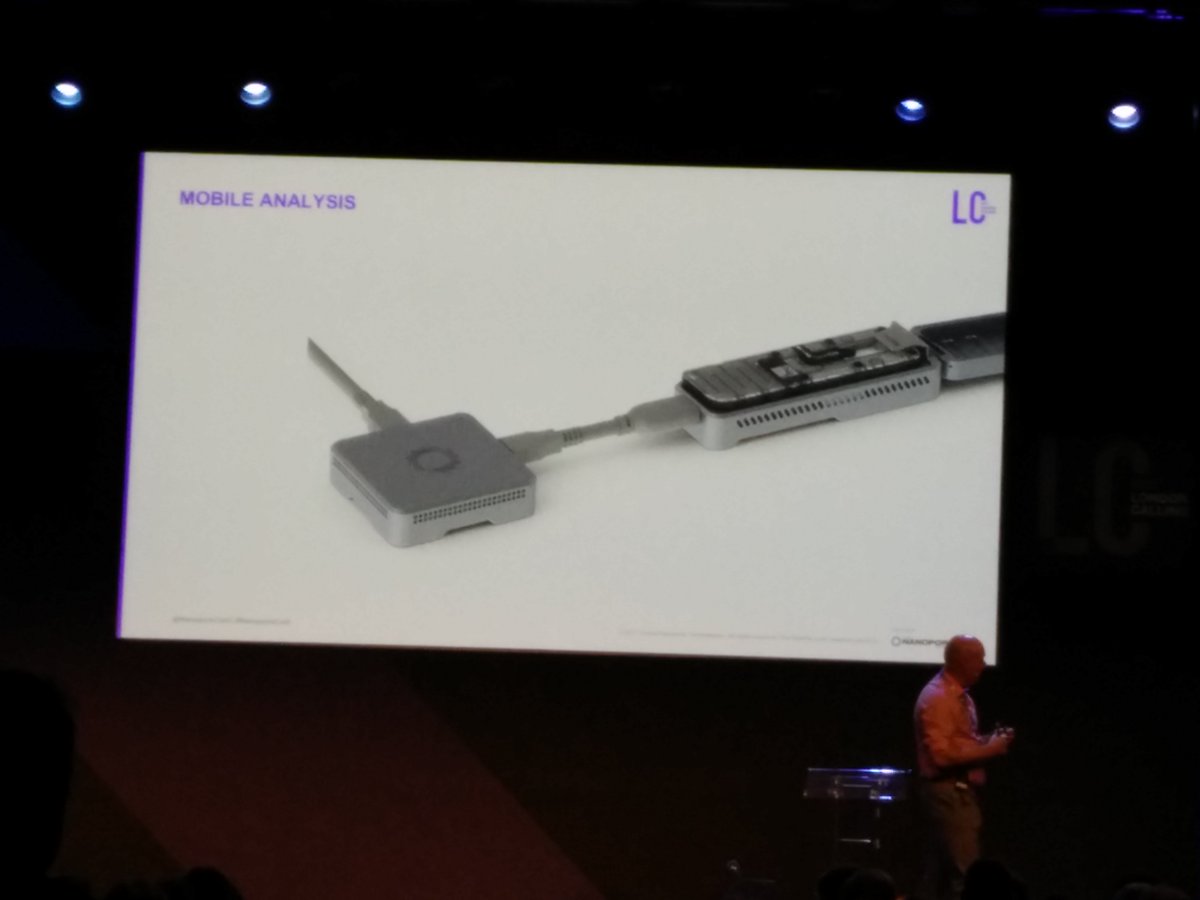
Perhaps the most notorious behind schedule ONT system is PromethION, which ONT hoped to have running about a year ago. Clive thanked early access (PEAP) participants for their patience and begged for a bit more. A few burn-in flowcells have gone out and users reported some were dead-on-arrival due to buffer leaking out. The need to invent a new flowcell manufacturing process was one of the many risks I flagged back in November. Many participants aren't even that far along; machines remain behind schedule on shipping. So there were no presentations of PromethION data by users at the conference, one of the many of my conference projections that did not come true. Clive vowed that getting PromethION launched this year is the company's number one goal. Brown also emphasized the message that PromethION should be able to exceed NovaSeq in total throughput while enabling more flexible workflows. It is his claim that PromethION will "turn the market around from short reads to long reads".
PromethION has evolved a bit further in design. Flowcells are now divided into four sector, each fully independent. Each sector has 750 pores, or about 1.5X that of a MinION flowcells, but requires only half the library load of a MinION flowcell. So each library aliquot for MinION should generate 3X the amount of data on PromethION. Two-fold multiplexing of channels is planned, with running life of four days. So if all this falls together, a standard library prep should generate 3X the amount of data by seeing more flowcells, and then some bump up from the longer runs. That increase is hard to compute, as outside ONT no one seems to see great performance towards the end of a run, so running twice as many days is not believed to yield anywhere near twice as much data. So conservatively two full PromethION flowcells, requiring the equivalent of 4 MinION flowcells of library, should generate in the field in excess of 120 gigabases of data, or about a human genome at 30-40X coverage.
Port geometries allow for using standard multichannel pipettors. Flowcells do still require priming, though the approach is a bit different and involves pushing buffer through the system from the waste port rather than pulling the bubble out through a priming port. Priming volumes are reduced, but wiping liquid off the flowcell seems to be required after each priming step. Flowcells include the expensive ASIC electronics; the "crumpet chip" design has proven difficult to achieve with the number of connectors required for PromethION (more on crumpet later).
GridION X5
GridION X5 boxes haven't hit the wild yet, so no data or experiences. Since these use MinION flowcells, they should perform like MinIONs. The first units should ship mid-May (15th). ONT believes they can build five machines per day, aiming to ship instruments 48 hours after shipping. Brown also promises that the on-board compute, augmented with FPGAs, will be able to keep up with all five flowcells even should ONT increase the sequencing speed to 1000 bases per second -- a velocity frequently mentioned but with no timelines.
I talked to a number of people who are seriously contemplating purchasing a GridION, but no one who had committed -- though I realized I never pinged someone I'm pretty sure is planning on one. Many who might consider one have already invested in the PromethION and remain optimistic that will come online in the near future. For both big boxes, ONT is touting them as enabling workgroups to run independent workflows asynchronously, much as you might treat a cold room or warm room as shared space.
Oxford reiterated their pricing models for GridION, emphasizing that they would prefer to sell all their gear on a "pay-as-you-go" basis but they threw in the capital purchase model for customers who have capital budgets which can't be easily converted to consumable budgets.
SmidgION and Flongle
One of the electrifying announcements last London Calling was the SmidgION concept for a sequencer that docked into iPhone. This was followed in October with the Flongle concept, an adapter to enable SmidgION flowcells to be run on MinION. SmidgION chips were promised to use the crumpet design. Between the smaller size and electronics separation, SmidgION offered an opportunity for extremely low cost consumables. Both Brown and Sanghera touted the opportunity to enable inexpensive diagnostics with Flongle. Nick Loman's talk later illustrated this point: hundreds of MinION flowcells lightly used in the Ebola project, since getting information rapidly was a priority and precluded batching. Indeed, Brown declared that ONT would seek regulatory approval for Flongle. Timelines weren't given for the appearances of these devices.
ONT is now projecting Flongle by the end of the year with SmidgION to follow sometime next year. Flongle/SmidgION flowcells will have either 128 or 256 channels (it isn't clear in the tweets or my memory if this represents two flowcell models or a design decision not yet made). With the problem of crumpet design solved on Flongle, Brown hinted heavily that ONT will be revisiting the concept on MinION, describing the integrated wet-dry flowcells as "something that seemed like a good idea at the time".
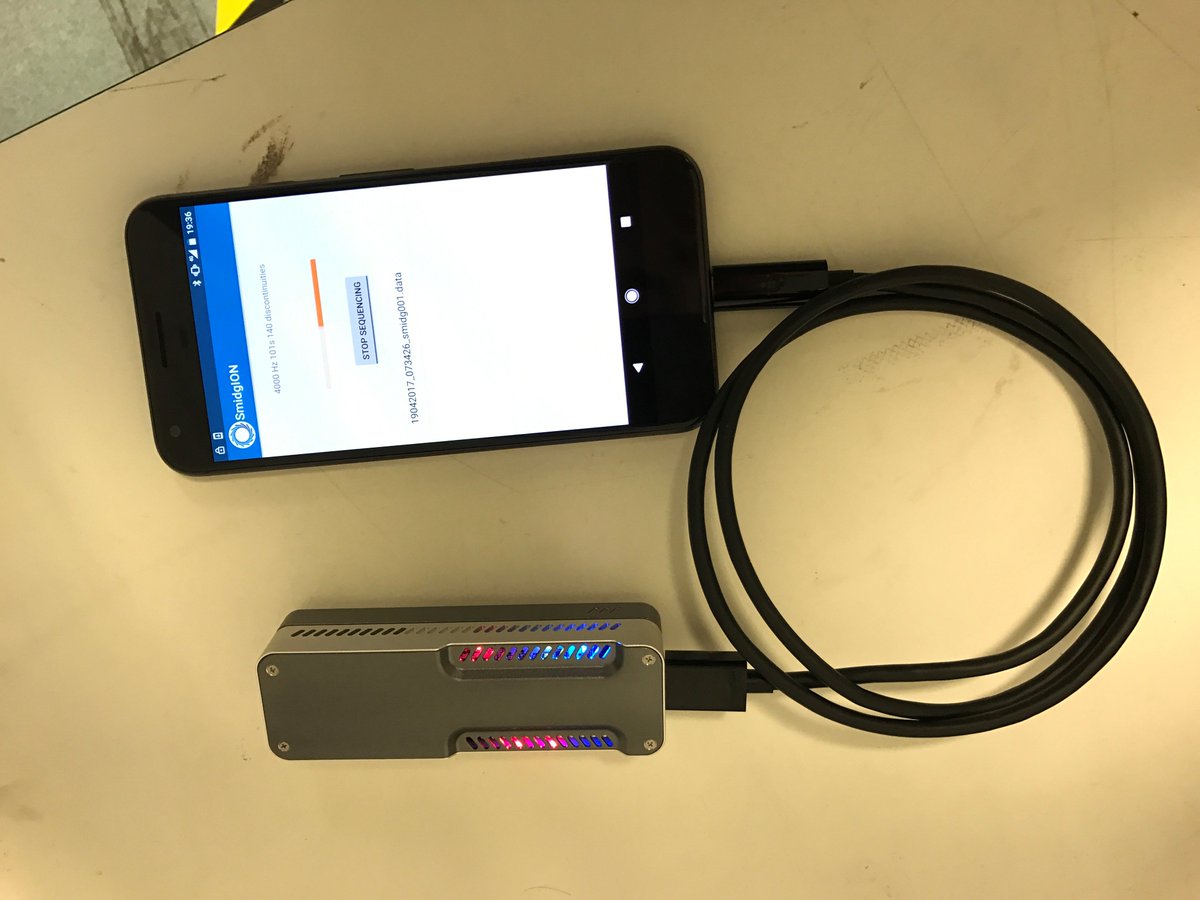
CEO Gordon Sanghera let me handle a SmidgION prototype. Designed to dock into a lightning port, it suffers the usual challenge with such schemes: the case which prevents me destroying my iPhone also blocks the SmidgION via steric hindrance. Of course, a lightning connector also excludes Android and Surface users. My iPad has a much less armored case and did accept the device. Someone else from ONT was showing a MinION running off an iPhone, connected via a lightning-to-USB cable. One issue with this is the MinION apparently has a voracious appetite for power, draining the iPhone in about two hours.
CEO Gordon Sanghera let me handle a SmidgION prototype. Designed to dock into a lightning port, it suffers the usual challenge with such schemes: the case which prevents me destroying my iPhone also blocks the SmidgION via steric hindrance. Of course, a lightning connector also excludes Android and Surface users. My iPad has a much less armored case and did accept the device. Someone else from ONT was showing a MinION running off an iPhone, connected via a lightning-to-USB cable. One issue with this is the MinION apparently has a voracious appetite for power, draining the iPhone in about two hours.
Brown also projected that Flongle would support the first appearance of solid-state nanopores. These might not be initially capable of sequencing but instead of "blob counting" -- detecting specific tags (much like Nabsys or Two Pore Guys).
VolTRAX
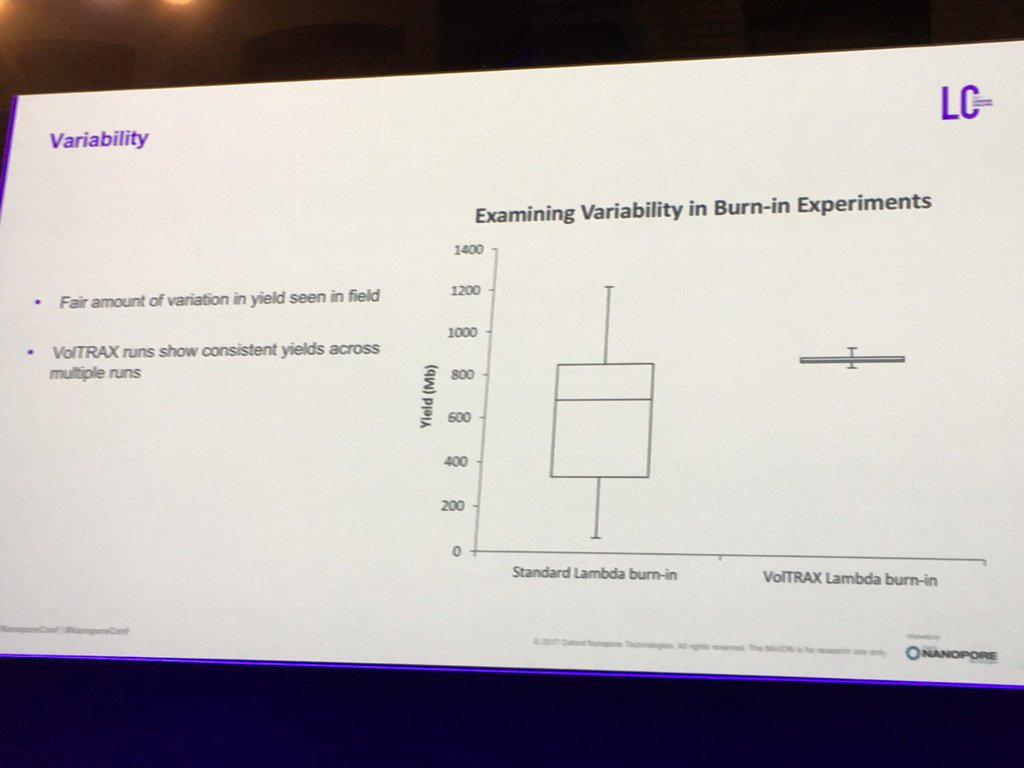
The VolTRAX library preparation device was much discussed and demonstrated, but access program participants haven't yet seen actual kits -- just colored liquid kits to familiarize users with operation and test out the controlling electronics. Apparently some of the early configuration kits suffered from leaks and buggy software, but a flurry of post-conference tweets show multiple users succeeding with newly updated software and kits.Oxford's initial kit offering for VolTRAX will be a version of the Rapid 1D transposase kit. That kit is theoretically so simple even I could run it, but in practice ONT showed that the VolTRAX prep (if it works, as one person commented) greatly compresses the variability in library quality. That "burn-in" kit will be followed with a "high-throughput" kit which uses the transposase method to fragment the DNA but then uses on-cartridge bead cleanups to improve the yield from the library. That is expected to be followed with a barcoding version of the same kit. For those who like one-stop-shopping, the VolTRAX transposase kits require no external reagents: the kit is claimed to be complete.
There was great enthusiasm both at the meeting and on Twitter to get other kits on the system. ONT wants to put all their library methods on, but there seems to be a friendly tussle for what kit will go after the first three. Oxford has the REPLI-g multiple displacement amplification kit running on VolTRAX, enabling genomic libraries from 1 nanogram or less of input. However, there was certainly a lot of interest in seeing the direct RNA or strand-switching cDNA preps go on VolTRAX.
ONT has also demonstrated internally loading VolTRAX with E.coli cells and performing DNA extraction and library preparation as an integrated workflow. Since VolTRAX has both magnetic bead capabilities and two-zone Peltier thermal control, even PCR should be possible on the device.
Not strictly VolTRAX, but a nifty protocol that ONT has demonstrated is to perform very limited PCR on a region of interest and then sequence both that product and the background genomic DNA together. In other words, the PCR templates are sequenced along with PCR products. This requires careful balancing, but when it works one can get both DNA copy number profiles as well as targeted mutation information from the same sample in a single workflow. The example application, presented by Dan Turner, was for simultaneously screening embryos for aneuploidies and specific mutations of concern.
By end-of-year, ONT projects that they will be shipping VolTRAX cartridges with lyophilized protocol-specific reagents already loaded. These are planned to be stable at ambient temperature.
ONT is also trying to move a simple spectrophotometry setup onto VolTRAX so that DNA purity and quantity can be assessed during a VolTRAX run.
Zumbador
The Zumbador field device for DNA extraction and library preparation continues in development, but "not quite there yet".Direct RNA & Strand-Switching cDNA
One theme both in Clive's talk and some of the other ONT talks is that there is a method to ONT having multiple protocols which seem to cover the same space: each protocol has different advantages and drawbacks. This is particularly apparent in the cDNA space: there is the new direct RNA kit and the strand-switching cDNA kit. Why choose cDNA if you can have your RNA direct? Well, the RNA motor runs at a much slower speed than the DNA motor, so the RNA kit has inferior throughput. But, if you wish to look at RNA modification, then the direct RNA kit is the way to go. Oxford is planning to roll out even more interactive guides to help users choose the best protocol for their experiment, with the guides pointing out these tradeoffs.Both of these protocols have been undergoing major tweaks. The existing direct RNA kit actually has a reverse transcription step, but this simply provides a DNA strand to support the RNA which is actually being read. That step is being dropped out, resulting in a much shorter protocol, from about 3 hours to around a half hour. The strand switching cDNA protocol has been tuned for much higher throughput, and can now deliver over a million long cDNA reads (sometimes approaching 10 million reads). There was a bunch of discussion at the meeting and online, without definite conclusion, as to how adequate such a procedure alone would be for quantifying message isoforms (as opposed to using short read data -- much of which won't capture isoforms -- to perform the quantification).
There is also a variant version of the cDNA protocol incorporating PCR, enabling working with smaller inputs. Dan Turner displayed data suggesting that quantification is not substantially distorted by this workflow.
ONT continues to work on protocols to target high-value niche problems. For example, they have developed a "semi-specific" cDNA protocol to target messages in which only one side of the message is known -- i.e. for targeting fusion transcripts.
Basecalling
Basecalling and accuracy continue to be a focus. The new 1D^2 kits should be shipping now, though there was certainly a lack of consistent message from the ONT staff as to what is different in the 1D^2 workflows. R9.5 flowcells are supposed to ship this week, which are promised to be compatible with all prior library kits. Some sort of tweak in the platform was mentioned to be in the works which is claimed to boost yield by 20-30%, but neither a precise timeframe nor the nature of the change were specified.Because only a few alpha sites have seen 1D^2, there wasn't much non-ONT data on these beyond what is in the human genome preprint (which Jared Simpson covered to a degree in his excellent talk). Oxford claims 1D^2 can hit 1% error rate, with individual reads in the low single digits. Brown reiterated his belief that there is "still plenty of headroom" for improving accuracy and that it will be a process of identifying a major error mode, developing an attack on that mode and then reassessing errors and repeating the process. He also mentioned that the current method for combining the information from the two strands in 1D^2 is old and sub-optimal,
Clive touched a bit on the challenge of calling weird DNA chemistries, mentioning that some fungi glycosylate their DNA. Some T bacteriophage also do this and much more to try to evade restriction systems. Want stranger? Some Streptomyces place phosphorothioate linkages in their DNA backbones, for reasons still be unveiled.
Brown also reiterated a favorite notion of his that different pores (or pore mutants) may have complementary error profiles and ONT might release future flowcells with such pores or mixed pore populations. He also mentioned a pore with two "reading heads" that interact with the DNA in two distinct regions abount 30 bases apart.
Also coming down the pike is the option for users to generate FASTA or FASTQ output in place of the unloved FAST5 files. ONT is moving their basecallers away from working with segmented "event" models to just processing raw electrical signals.
One of the more "out there" items in Brown's talk is an idea of training networks to simply convert raw signal into final results (such as species calling from rRNA data) with no explicit intermediate basecalling. He described this as a pet project which he plans to spend time on. The scheme might be implemented, at least in part, on FPGA devices including the dongle.
MinKNOW
In part due to GridION X5 and PromethION, ONT is rolling out a new version of the MinKNOW control software. While the software was not available for viewing (I think -- didn't look hard), screenshots showed up in multiple talks. The new software is designed to better support multiple flowcells on the same machine, allowing users to group flowcells and then view aggregate statistics for these groupings. The new software is also intended to greatly simplify the interface (Brown cited an iPhone-like intuitive feel as the target), with easier protocol selection. The software will also give more feedback to users as to the quality of a run over time, presenting a chart of how key categories of pores (bad pores, actively sequencing pores, pores waiting for a DNA to work on) changes during the run.A big improvement for user friendliness will be renaming "Platform QC" to "Check My Flowcell", with a much simplified output format. Flowcells will be scored as Red (bad), Yellow (usable but below warranty) or Green (above warranty) -- and ONT did design the displays so that the color reinforces but is not necessary (hence friendly to the color-blind).
Another feature in development, though it wasn't clear when it would hit, is for the software to track the amount of use each channel has seen. This is important for flowcells that are washed and reused: the electrochemical gradient shifts over time and performance will suffer if this shift isn't accounted for.
An even more radical idea that isn't on track, apparently due to mixed opinions at ONT, is to dynamically change which pores are being used at any given time. At the beginning of a run, MinKNOW groups ranks the pores for each channel, creating four groups of pores. Typically the first two groups will be very robust in number and group 3 will be a bit less and group 4 (particularly for aged flowcells) downright anemic. During the course of a run, first group 1 is used and then at some point it is swapped for group 2 and later group 3 takes its turn (group 4 apparently sometimes is never used; I've certainly seen this behavior in our runs). But during the period a group is active, pores do die, reducing the yield. If you (or your patient!) is impatient, then it would be valuable to dynamically replace a newly dead pore with another pore from the same channel, if one is available. The catch is that the use history of the pore determines the required voltage: even unused pores leak the electrochemical mediator slowly and pores in use leak it more quickly. So complicated on the software side, but potentially a winning strategy to even more heavily front-loading data collection.
Targeted Quantification
ONT presented strategies for tag-based quantification of both nucleic acids and proteins of interest. For the nucleic acid case, Cas9 with custom guide RNAs (which Oxford will have tools to enable design, and may serve as a vendor for the guides) with specific tags on the Cas9 proteins. As noted above, these might not be actually sequenced but "blob counted" on solid state pores. Cas9 can also be used to enrich specific genomic targets. This product should launch this year, with the tentative name "Cas me if you can". ONT personnel dodged any questions about the ongoing Cas9 patent litigation.For proteins, ONT is focused on a sandwich immunoassay scheme, in which one antibody is linked to a DNA probe to be detected. Well, at least their slides showed a sandwich approach; presumably a zoo of DNA-linked anti-beast antibodies (anti-mouse, anti-goat, anti-camel, anti-jackalope -- you get the idea) could be used for multiplexing.
After the conference ended, a couple of ONT folks sought me out to describe a vision of "omniome" -- profiling DNA, RNA and protein from the same sample on the same flowcell. Such a protocol would branch into three branches, but these would be brought back for the final sequencing. Trying to get that on VolTRAX wasn't mentioned, but that technology is capable of splitting liquids and could make such a trident-scheme practical at scale.
Improving Sensitivity and Consistency
Brown in his talk and several other speakers from ONT reiterated the belief that much of the yield variability seen by users is due to suboptimal DNA preps or DNA input into library preparation. The new displays in MinKNOW should give more feedback to users. Interestingly, Mick Watson mentioned that he has had repeated 1D transposon kit failures on his sheep rumen microbiome samples and has gone to just ligation preps. We just had a similar failure on a HMW sample with excellent absorbance statistics (A260/A280 & A260/A230).One user reported a 12 gigabase flowcell which an unfortunate informatics problem (probably a full SSD -- been there, done that!) resulted in tossing about another 4 gigabases -- so a 16 gigabase fish tale (well, actually a plant tale). ONT has launched a new sequencing yield contest, with free flowcells to those who can beat set targets. Perhaps they should also have a bounty program for transposase-recalcitrant DNAs.
ONT is looking at ways to improve the sensitivity of the system. According to a plot shown in the workshop, optimal ligation libraries are achieved with 200 femtomoles of input DNA, which is 1 microgram of 8 kilobase fragments. Performance drops slowly with lower amounts, until it falls off a cliff around 10% of that amount. ONT would like to shift this curve to the left for understandable reasons. Want to load 200 fmol of 800kb DNA? That will be 100 micrograms please -- an absurd amount. Clearly this isn't needed; if such a 200fmol load generates 1 million reads, then only about 1 in 100,000 of the input molecules actually goes through a pore (check my math!).
They showed an image of the membrane architecture with loading beads on it; the beads were tiny compared to the membrane field and many were clustered along the perimeter. Since the pore is expected to be the middle of that structure (not clear what drives it there), those beads aren't effectively delivering DNA to the target. While the mechanisms for driving DNA to the pore weren't described, ONT apparently has multiple ideas for improving this aspect.
Genome Foundry, Metrichor
Brown mentioned that their foray into gene synthesis, Genome Foundry, is expected to have news later this year.Clive also emphasized that Metrichor isn't just a software platform, but a separate spin-out company that plans to commercialize a range of measurement applications. The EPI2ME software will soon have a workflow builder allowing users to implement their own workflows.
'Til Next Post
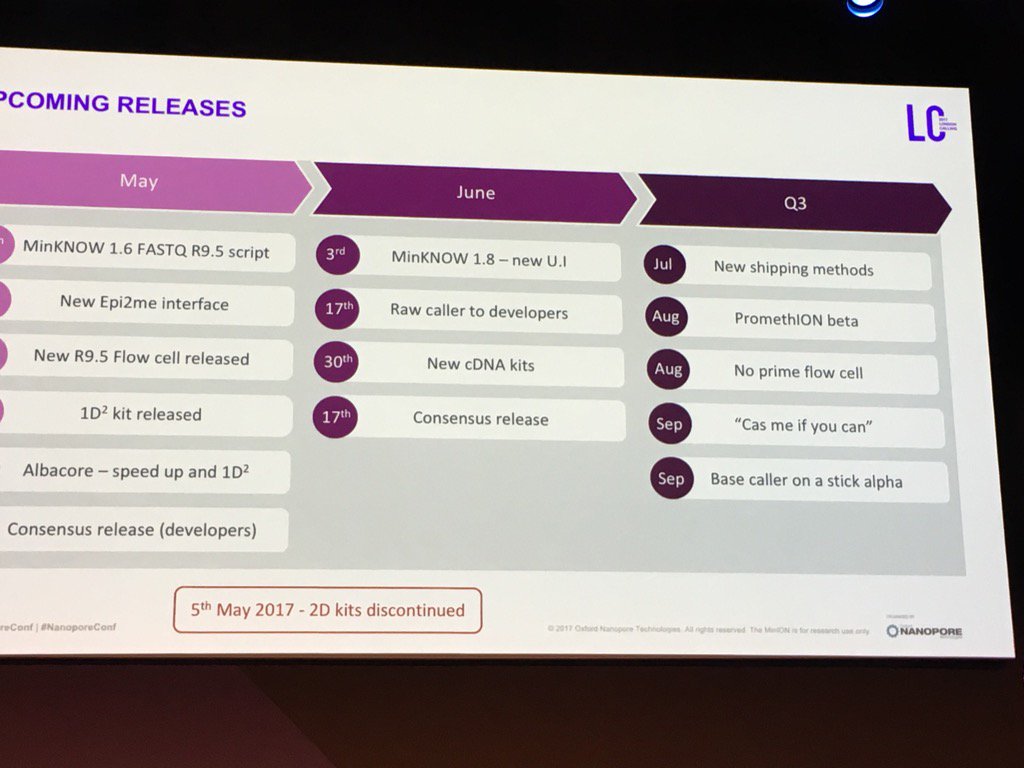
In the coming days I'll cover a bit of the user science at the conference and some other themes -- not sure how long I'll milk this. I didn't frame this shot quite right (cut off the days in May), but below is the projected dates for many of the items I have discussed.

10 comments:
Milk it keith...milk it for all its worth!
I'm with James - keep it coming!
Would be interested in your editorial on the platform progression - in particular how the GridION would stack up against the usual band of alternatives.
Seems to me that they are currently not equipped for full commercialization at the moment, and are not hiring people to support wide adoption...
Will your flowcells expire before you get all 48 replaced on the promethion? Good to see the ONT PR machine in full swing, but for a system that's supposed to be ultra-high throughput there seem to be a lot of moving parts. The UE on the promethion seems particularly cumbersome. We all used to complain about oiling a prism on the Illumina GA, now we have to worry about replacing 48 flowcells on a promethion every 10 days? Compare that to loading a NovaSeq or a Sequel and at least in a clinical environment there are clear winners and losers here.
Promethion will be a beta sequencer until late 2018 (ONT says early 2018 but they're always late). I wouldn't expect the users to expect an easy to use, robust system until they are out of beta. These things take time and so does building an organization with the proper commercial infrastructure. I suspect by 2019-2020 ONT will have a more robust promethion with a large commercial support team that allows them to reach the broader market. The question is where will Illumina/Pacbio be by that time.
"The UE on the promethion seems particularly cumbersome. We all used to complain about oiling a prism on the Illumina GA, now we have to worry about replacing 48 flowcells on a promethion every 10 days?"
Surely plugging something in 48 times takes about 2mins. How lazy are you? Also surely it can sit empty when not in use, as can a sequel or any other machine.
NB:\\ I have no association with any seq company I just think you are lazy!
Anonymous #1, another great point or even where Genia or someone whose still in stealth mode will be by then. It's going to get interesting.
Anonymous #2, You think it's lazy to criticize a "system that will beat the novaseq" for having a horrible user experience? Whose output is predicated on the use of 48 "flowcells" to achieve it's goal of defeating the highest output instrument on the market? You know what, 48 Illumina flowcells on a GA IIx also have more output than a NovaSeq.
The only thing here that is lazy is your argument.
It would be interesting for the community to hear some of your inputs on fundamental aspects of the ONT technology. For instance, do you see it as an alternative to PCR? Where is a market need for same day sequencing (PCR being gold standard) or is MinION more of a cheaper, long-read tool (as an alternative of Pac-Bio) where the portability is an add-on advantage? What is the problem space in the nether world between PCR & high-throughput Illumina sequencing?
"The only thing here that is lazy is your argument."
I wasn't making an argument, I was stating a fact. Your original objection was that you didn't want to load 42 flowcells; I was merely pointing out if that's your only issue with Promethium then you are lazy as it would take about 2mins.
You don't need to run all the flowcells. You can run any number, or none. They can even be loaded offline and plugged in. Run, then stored offline. The cost is the same. Yes 12Terbases is from all 48 over 12 hours.
Have you seen the size of a novaseq flowcell, you could go sledging on it, or use it to shovel snow in winter time. It's about 48 times bigger than GAII flowcell.
Downplaying the availability and the commercial capability of the company, Andy up playing pacbio and other potential rivals is just desperate naysaying. No rational basis at all other than I'll-wishing.
Anonymous #1, another great point or even where Genia or someone whose still in stealth mode will be by then. It's going to get interesting.
Please click to play,if you wanna join casino online. Thank you
gclub
จีคลับ
gclub
Gclub
Post a Comment